GESV pipeline
The GESV (Genome-edited Sequence Variants) pipeline assembles sequence variants (SVs) from amplicon data and compares them to a reference sequence to identify edit types and frequencies. By condensing raw amplicon data into distinct SVs while preserving their abundance information, GESV significantly streamlines amplicon sequencing–based genotyping of gene-edited organisms.
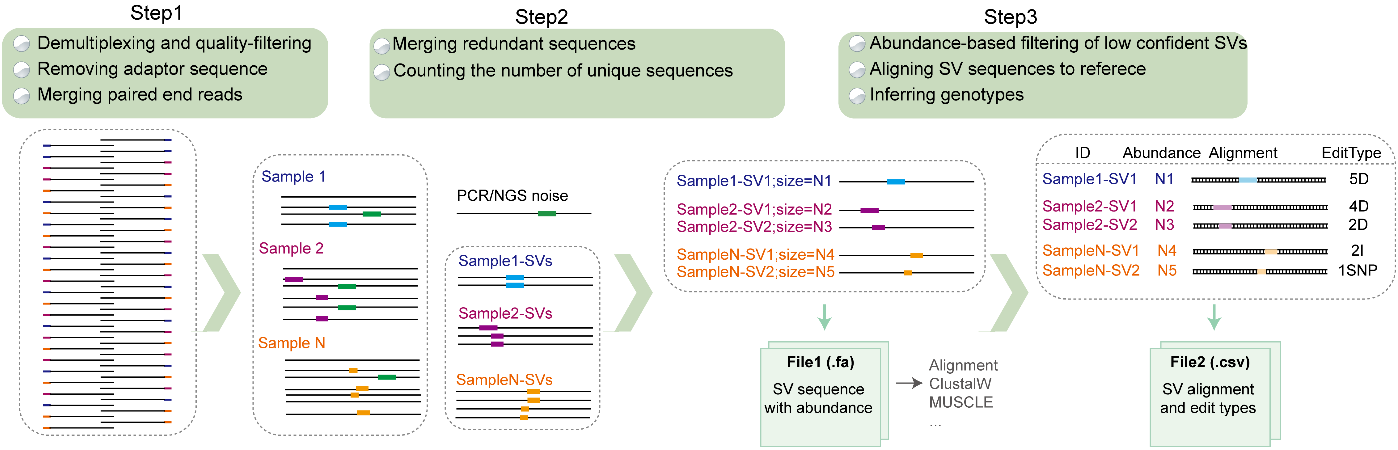
Schematic workflow of GESV analysis
Run GESV pipeline
To run the GESV analysis online, please provide sample information and upload the ZIP file containing raw FASTQ data.
Download offline gui apllication to run GESV pipeline locally, GESV-local .
More information about GESV pipeline can be found in the Hu_et_al_GESV_2025 GitHub repository.